Genomics
Knowledge about the clonal evolution of each tumor can inform driver-alteration discovery by pointing out initiating genetic events as well as events that contribute to the selective advantage of proliferative and potentially drug-resistant tumor subclones.
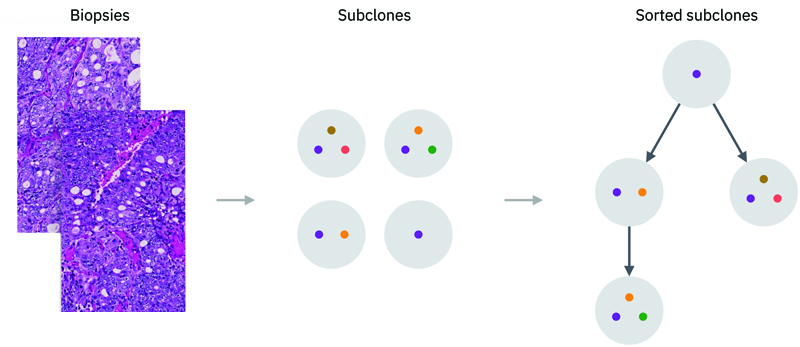
A necessary building block for the reconstruction of clonal evolution from tumor profiles is the estimation of the cellular composition of each tumor subclone cellularity. These, in turn, are based on estimates of the relative abundance frequency of subclone-specific genetic alterations in tumor biopsies.
Estimating the frequency of genetic alterations is complicated by the high genomic instability that characterizes many tumor types. Chimaera is an algorithm that allows an accurate detection and estimation of subclone frequencies using WES and WGS data from multi-area biopsies that enables the subclones to be sorted in an evolutionary tree structure.
Reference
“Inferring clonal composition from multiple tumor biopsies,”
Matteo Manica et al., arXiv:1701.07940, 2017.
Questions?
Ask the expert

Matteo Manica
IBM Research Scientist
Funding
This research is funded by the PrECISE EU project
